Our projects aim at understanding the bases of transcription regulation in the context of chromatin. We use various model organisms from yeast to mammals and have a focus on the role of secondary DNA structures in transcription control. We develop 4 main topics.
- Role of G-quadruplex secondary DNA structures to promoter and other regulatory module activities through nucleosome exclusion
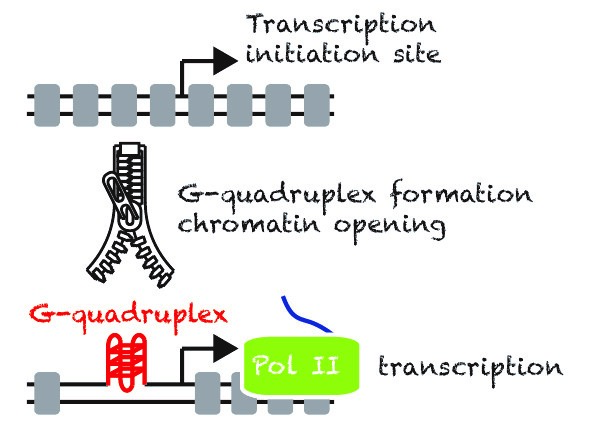
Despite many efforts in the past, the scientific community still does not understand what defines a promoter in eukaryotic cells. While most of the proposed sequence determinants of promoters are poorly conserved at the genome-wide level or during evolution. We propose that in mammalian cells secondary DNA structures called G-quadruplexes (G4s) are Nucleosome Exclusion Modules required for promoter activity. Thus, beyond the primary DNA sequence, the structures may play an essential role to sustain a robust promoter activity, more specifically at highly and widely expressed genes.
We have developed a technique, G4access1, to map G4s genome-wide for G4 forming sequences in the context of chromatin. This allowed us to show that G4 formation is mutually exclusive with DNA methylation, including at imprinting loci. Using G4access and by mutating G4s in promoter context combined to single-cell live-imaging, our recent work has shown that G4s directly act as promoter elements2. We are now working at better understanding how G4s control regulatory elements such as insulators and enhancers and how G4 regulators influence gene expression, chromatin opening, DNA supercoiling and stability.
Main collaborations: Jean-Louis Mergny (Ecole Polytechnique, Orsay, France), Salvatore Spicuglia (TAGC, Marseille), Kateryna Makova (Penn state University), Daan Nooordemer (I2BC, Orsay).
- Molecular bases of chromatinopathies
Chromatinopathies are a group of human diseases that involve mutations in chromatin-regulating proteins and enzymes. These mutations are often acquired de novo and many are monogenic. For example, some complexes involved in histone methylation or acetylation can lead to Kabuki or Rubinstein-Taybi syndromes which are associated with neurological alterations and other symptoms such as visio-spatial defects, heart malformation and immune defects3-5. In close collaboration with clinicians of the Montpellier hospital and other European labs, part of a MSCA Chrom_rare consortium, we develop human iPS and zebrafish models to study these diseases, focusing on the Kabuki syndrome. This syndrome involves monogenic mutations of the KMT2D and KDM6A genes. We also investigate other Kabuki-related haplo-insufficient models such as CDK13 and other cases altering secondary DNA structures, such as G-quadruplexes. We monitor their specific chromatin and gene expression alterations and explore treatment options in animal models (Kabuki-related zebrafishes) with our collaborators.
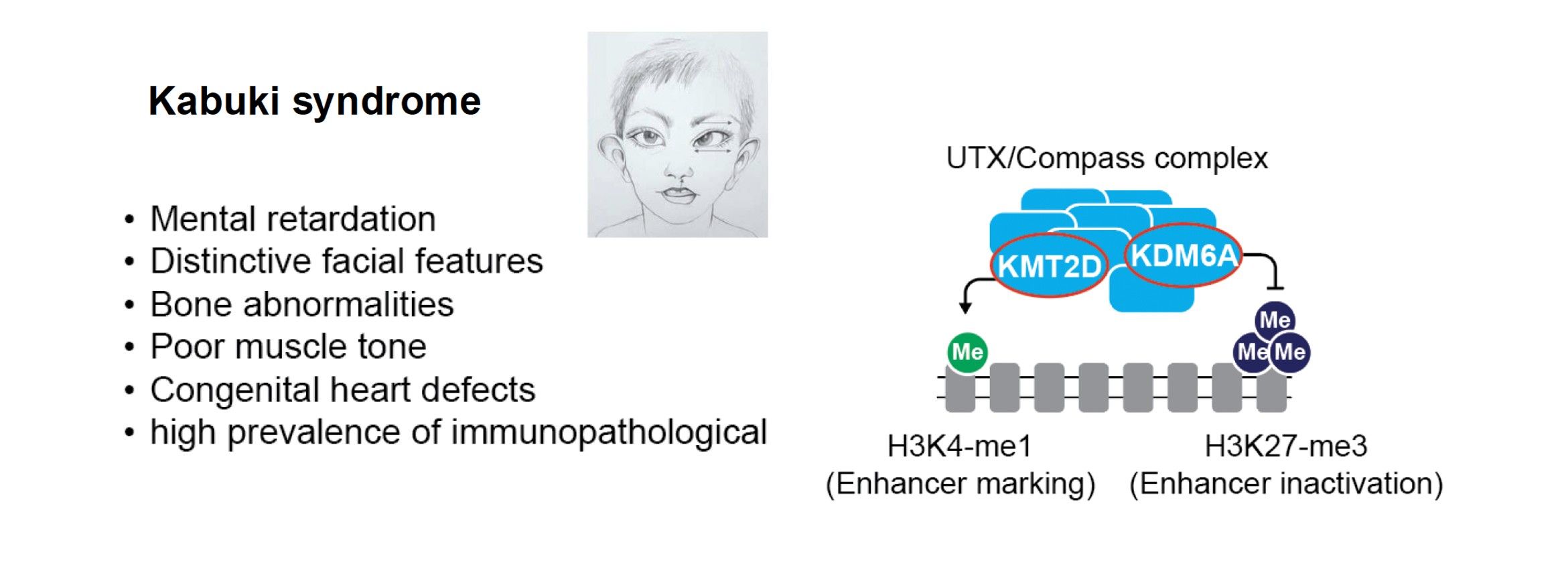
Main collaborations: David Genevieve (CHU, Montpellier), Farida Djouad (IRMB, Montpellier), Aleksandra Pekowska (Nencki Institute, Warsaw, Poland), Siddharth Banka (Un. Manchester, UK).
- Mechanotransduction roles in 3D genome organization and gene expression programs (program lead by C. Esnault)
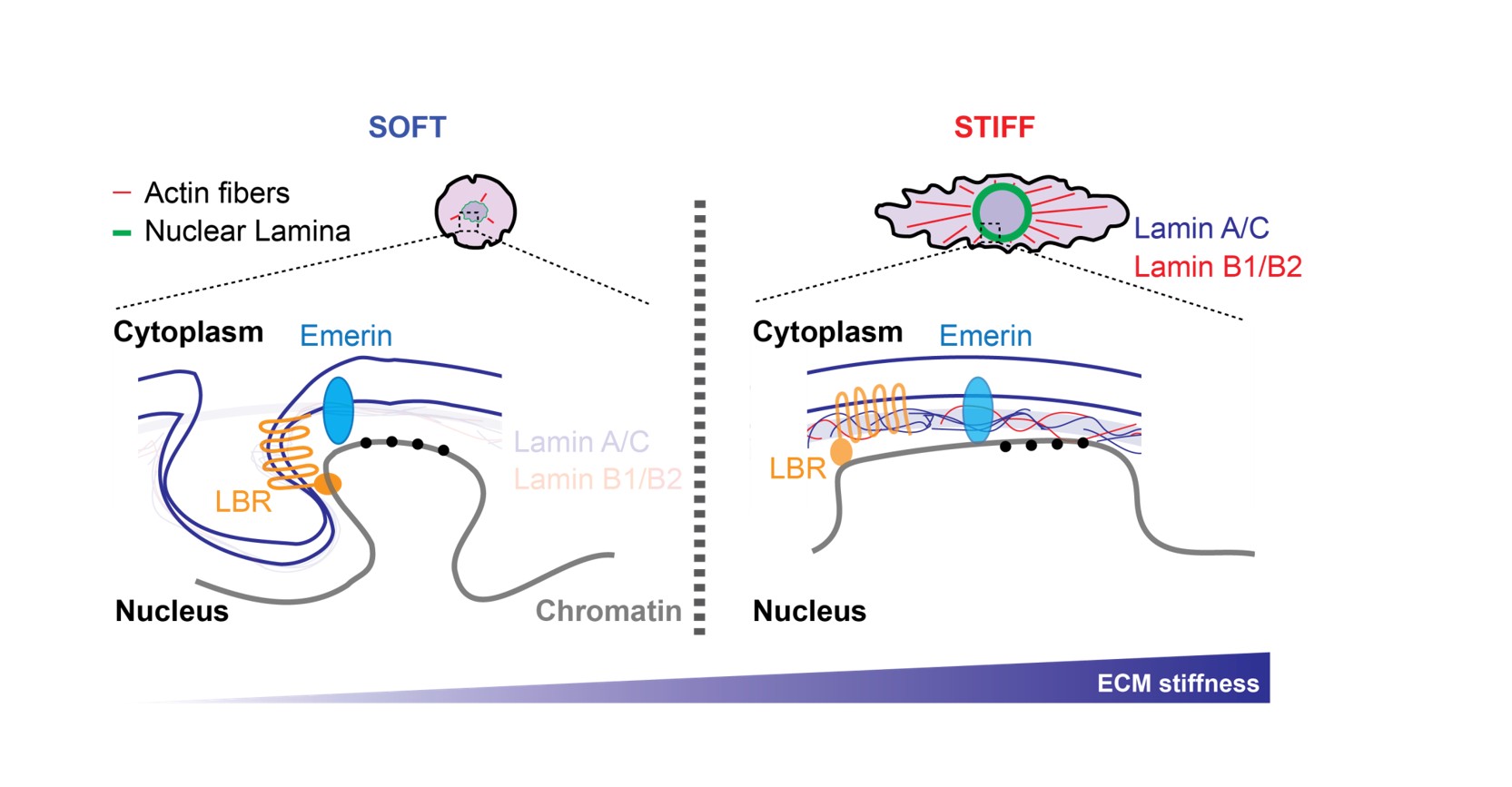
Environmental cues, including growth factors, hormones and nutrients are essential for regulating cell behaviour. Alongside of these biochemical signals, cells also respond to physical constraints, such as external forces: stretch, compression and extracellular matrix (ECM) stiffness. ECM elasticity for instance, is of critical physiological importance (i) it regulates epithelial and mesenchymal stem cell differentiations, (ii) promotes fibroblasts transformation into cancer-associated fibroblasts (CAFs) and (iii) supports breast cancer progression. These processes are underpinned by changes in transcription and epigenetic programming. Using Omics approaches, we investigate how nuclear membrane and cytoskeleton dynamics shape the 3D genome organization and modulate gene transcription. Our most recent work has emphasized Lamin and PRC2 Polycomb complex interplay during the transition from soft to stiff matrices.
Main collaborations : Kerstin Bistricky (CBI, Toulouse), Manoel Manghi (LPT, Toulouse)
- Régulation de la transcription
Our lab has been interested in the control of transcription by the RNA polymerase II CTD, showing the importance of this domain and that of its Tyr residues in termination control6,7. We also showed that the dynamic nature of enhancers during differentiation can condition isoform selection during mouse T-cell differentiation8. In a collaborative frame, we now investigate how transcription proceeds in the quiescent state (G0), using yeast as a model organism.
Main collaboration : J. Wu, ICGB, Bordeaux
References
1 Esnault, C. et al. G4access identifies G-quadruplexes and their associations with open chromatin and imprinting control regions. Nat Genet, doi:10.1038/s41588-023-01437-4 (2023).
2 Esnault, C. et al. G-quadruplexes are promoter elements controlling nucleosome exclusion and RNA polymerase II pausing. Nat Genet 57, 1981-1993, doi:10.1038/s41588-025-02263-6 (2025).
3 Aref-Eshghi, E. et al. Evaluation of DNA Methylation Episignatures for Diagnosis and Phenotype Correlations in 42 Mendelian Neurodevelopmental Disorders. Am J Hum Genet 106, 356-370, doi:10.1016/j.ajhg.2020.01.019 (2020).
4 Comel, M. et al. Abnormal Immune Profile in Individuals with Kabuki Syndrome. J Clin Immunol 45, 7, doi:10.1007/s10875-024-01796-5 (2024).
5 Margot, H. et al. Immunopathological manifestations in Kabuki syndrome: a registry study of 177 individuals. Genet Med 22, 181-188, doi:10.1038/s41436-019-0623-x (2020).
6 Shah, N. et al. Tyrosine-1 of RNA Polymerase II CTD Controls Global Termination of Gene Transcription in Mammals. Mol Cell 69, 48-61 e46, doi:10.1016/j.molcel.2017.12.009 (2018).
7 Yahia, Y. et al. RNA polymerase II CTD is dispensable for transcription and required for termination in human cells. EMBO Rep, e56150, doi:10.15252/embr.202256150 (2023).
8 Maqbool, M. A. et al. Alternative Enhancer Usage and Targeted Polycomb Marking Hallmark Promoter Choice during T Cell Differentiation. Cell Rep 32, 108048, doi:10.1016/j.celrep.2020.108048 (2020).



