We are interested in the role of chromatin in the epigenetic transmission of cellular phenotypes. We have two main projects to study the mechanisms of chromatin structure maintenance after genome replication and cell division:
1.Chromatin dynamics during DNA replication
(ERC- consolidator grant NChIP 647618; 2015- 2020)
Chromatin assembly is a fundamental cellular process necessary for the maintenance of genome integrity and transcriptional programs. Understanding the effect of DNA replication on histone protein dynamics is also a prerequisite for understanding the role of chromatin in epigenetic inheritance. Epigenetic phenomena are thought to influence cellular differentiation and cancer formation, as well as the impact of environmental factors on early development and later predispositions to disease. While epigenetic inheritance of chromatin components is, in theory, accepted as the driver of such phenomena, chromatin state inheritance per se has only been demonstrated for a few specific cases. Not much is known about histone “inheritance” beyond the facts that bulk maternal histones distribute equally among the daughter strands and are diluted two-fold after replication with newly synthesized “unmarked” histones, and that the majority of H3/H4 tetramers do not split before reassembly. We have shown previously that maternal nucleosomes stay on average within 400bp of their original binding site, implying that any potentially heritable chromatin encoded information, has to be inherited in ~1kb blocs, as smaller nucleosome domains would rapidly be diluted by new nucleosomes.
We are developing high throughput techniques (such as NChAP-Nascent Avidin Chromatin Pulldown; Figure 1) for directly measuring movements of histones and chromatin regulators during genomic replication in S.cerevisiae to determine, how chromatin states survive the perturbations associated with replication. This will allow us to assess locus specific differences in the spread of maternal nucleosomes after replication, the effects of leading and lagging strand replication on nucleosome positioning and maternal nucleosome distribution, the renewal dynamics of posttranslational histone marks and chromatin binding proteins, and the kinetics of chromatin footprint re-establishment and gene (re)activation.
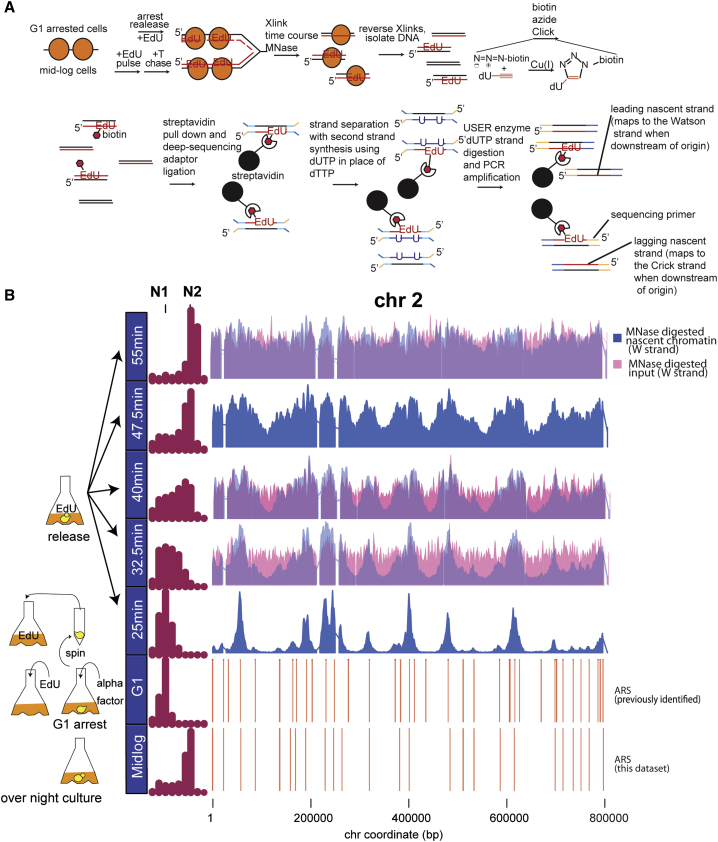
Figure 1: Nascent Chromatin Avidin Pull-Down
(A) Diagram of nascent chromatin avidin pull-down (NChAP). For synchronized cells, after arrest in G1, cells are released into fresh media in the presence of EdU and aliquots are fixed at regular time intervals. In asynchronous populations, cells are pulsed with EdU, followed by a thymidine (T) chase. Chromatin is digested with MNase, and the isolated DNA fragments are subject to a click reaction that adds biotin to the incorporated EdU. Biotinylated DNA is purified with streptavidin-conjugated magnetic beads, and NGS libraries are constructed on DNA fragments attached to the beads. cDNA strands are separated with primer extension in the presence of dUTP. The dUTP-containing strand is then digested with USER enzymes prior to PCR. This ensures that only nascent strands are sequenced.
(B) Density distribution of DNA content measured by flow cytometry before arrest (mid-log) in G1 and at indicated times after release from G1 arrest (left panel). Nascent chromatin Watson (W) strand read distribution on chromosome 2 at indicated times after release (blue bars) and total chromatin input are shown (total MNase-digested chromatin isolated prior to the click reaction; 32.5-, 40-, and 55-min time points, pink bars). Replication origins (ARS) are shown in the two bottom rows: ARS from this study (first) and previously documented ARS (second) are shown. Read counts were grouped in 400-bp bins and first normalized to the genome average read count and then to the highest peak value in each chromosome.
2.Asymmetric Segregation of Chromatin components
(ANR GENCHROSEG; 2014-2018)
Asymmetric cell division is a prerequisite for cellular differentiation and stem cell maintenance. Phenotypic transformation during differentiation is a poorly understood epigenetic phenomenon, in which chromatin, as a transcriptional regulator, theoretically plays a role. The underlying assumption that chromatin components segregate asymmetrically in asymmetric divisions has however not been systematically tested. Budding yeast also undergoes asymmetric divisions producing mother and daughter cells. The mother can generate 20-30 daughters during its replicative lifespan, and most of the factors that determine the phenotypic identity of the mother cell are unknown. The high conservation of chromatin components among eukaryotes and the availability of powerful genetic tools make S. cerevisiae an ideal system for identifying chromatin components involved in
asymmetric division and elucidating the underlying molecular mechanisms, using whole genome and single cell approaches (Figure 2).
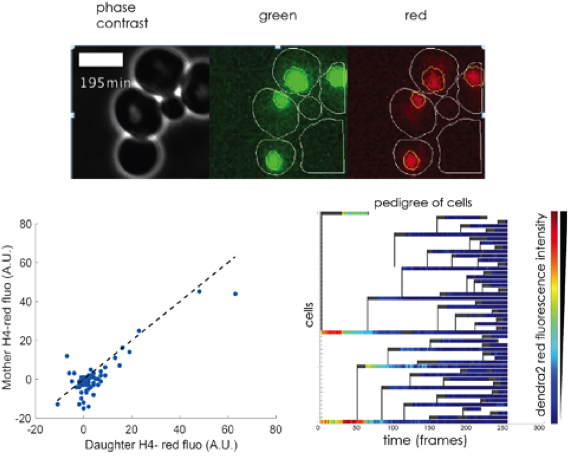
Figure 2
HHF2dendra2 lineage in a microfluidic chamber. Dendra2 is a photoconvertible fluorescent protein that we fused to histone H4 (HHF2). The photo conversion feature allow us to differentiate between already synthesized maternal proteins, which will fluoresce in the red spectrum after a blue laser light pulse, and the green fluorescent proteins synthesized after the blue light pulse. Microfluidic chambers make it easier to automatically identify mother and daughter pairs. Top: frame from a film of dividing cells (photos taken every 3 min). Time 0 is right after the photo-conversion. Bottom: The red signal is divided equally between the mother and the daughter cells until it is reduced to background levels in 2 to 3 generations, as shown in the mother daughter red fluorescence scatter plot on the left and the pedigree heat map on the right.