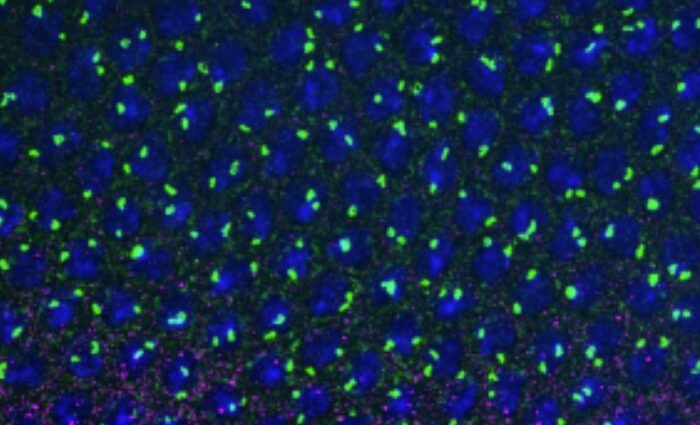
The helicase senataxin (SETX) has been the object of intense investigation since the discovery of its involvement in two unrelated neurodegenerative diseases, almost two decades ago. Recessive mutations in the gene encoding SETX lead to ataxia with oculomotor apraxia, while dominant mutations are associated with a juvenile form of amyotrophic lateral sclerosis. SETX has been involved in different facets of RNA metabolism as well as in the maintenance of genome integrity by controlling the levels of particular structures named R-loops. These structures form in certain contexts during transcription when the nascent RNA anneals with the DNA template, thus leaving the non-template DNA strand unpaired. Unscheduled persistence of R-loops can lead to the accumulation of DNA damage.
However, the precise role of SETX in RNA metabolism and the control of R-loops have remained unclear.
In this study, we have purified the helicase domain of SETX and provided the first biochemical characterization of SETX catalytic activities. Importantly, using various in vitro assays, we have demonstrated that SETX can directly induce the termination of transcription by the RNA polymerase II and that it can dismantle synthetic R-loops. In addition, we have analysed two SETX variants harbouring disease-associated mutations and shed light into the effect of such mutations on SETX folding and biochemical properties.
Altogether, these results broaden our understanding of SETX function in gene expression and the maintenance of genome integrity and provide clues and tools to elucidate the molecular basis of SETX-associated neurodegenerative diseases.
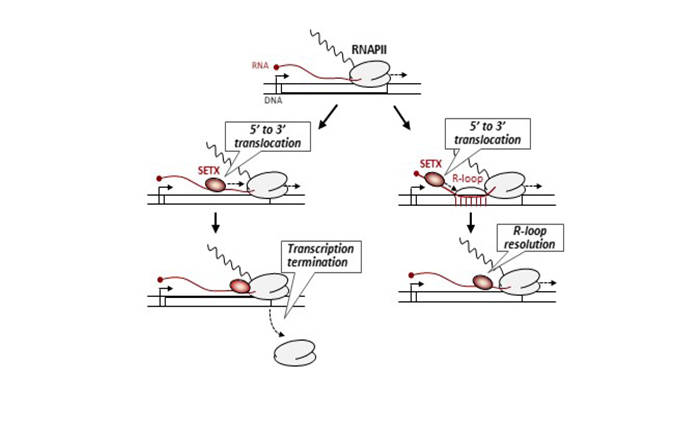
Figure legend: Schematic model of the mechanism by which SETX promotes transcription termination and R-loop resolution. RNAPIIs may pause/stall during transcription for instance upon reading specific DNA sequences or upon encountering other DNA-bound proteins or damaged template. SETX translocates on the single-stranded nascent RNA with a 5′-3′ polarity to dislodge RNAPII and its associated transcript. R-loops formed during transcription most often possess a 5′ RNA overhang. SETX can efficiently resolve this kind of structure by translocating in the 5′-3′ direction on single-stranded RNA to disrupt the RNA:DNA hybrid within the R-loop and release the RNA moiety.
Reference : Human senataxin is a bona fide R-loop resolving enzyme and transcription termination factor (2023). Zdenka Hasanova, Veronika Klapstova, Odil Porrua*, Richard Stefl*, Marek Sebesta*. Nucleic Acids Research, gkad092, https://doi.org/10.1093/nar/gkad092