Our group has a long-standing interest in inflammation and tumorigenesis. We have a strong record in studying Tumor Necrosis Factor (TNF) family members and pioneered the characterization of the biological role of the TNF-like ligand A Proliferation Inducing Ligand (APRIL), a protein that was originally identified by the PI. In particular, we discovered that APRIL acts as a pleiotropic cytokine that dampens inflammation and promotes B cell malignancies and colorectal cancer (CRC). Our expertise in mouse models for intestinal inflammation and carcinogenesis initiated several new projects and collaborations, as illustrated by our recent publication in the Journal of Clinical Investigation (doi: 10.1172/JCI131517) describing a yet unappreciated bifunctional role of cyclin A2 in CRC. This work has been highlighted by CNRS news in January (https://insb.cnrs.fr/index.php/fr/news-list). More recently we started to focus on a yet understudied aspect of specific posttranslational modifications in CRC and inflammatory diseases.
In 2021, the team previously led by Urszula Hibner, working on inflammatory liver disease and hepatocellular carcinoma (HCC), associated to our group, which will further strengthen our expertise and allow us to extend our research to different gastrointestinal tumor type. The team now led by Damien Grégoire investigate on oncogenic cooperation in hepatic carcinogenesis, focusing on functional consequences of defined genetic composition the tumor. The inter-tumoral heterogeneity is questioned by analyzing initiation and progression of tumors driven by different oncogenic alterations, combined with the study of the cross talk between tumors and the stromal environment.
Colorectal cancer (CRC) remains a major public health issue requiring the identification of new biomarkers and therapeutic targets. Together with the Mamessier team (CRCM Marseille) we combined relevant mouse models and a meta-analysis of transcriptome data of CRC to evaluate the role of cyclin A2 in CRC. Though high cyclin A2 levels have been frequently associated with bad prognosis in cancer patients, this new study displays that high levels of CCNA2 (the mRNA coding for cyclin A2) may represent a favorable prognostic marker in CRC.
Colorectal cancer (CRC) is one of the leading causes of malignancy-related death worldwide with increasing incidence. Disease status of patients can be classified according to histologic tumor grade (differentiation) and anatomic extent of disease (i.e., (TNM, tumors/nodes/metastases, stages I–IV), which describes tumor growth into the wall of rectum or colon (stage II), involvement of regional lymph nodes (stage III) and metastatic spread to other organs (stage IV). A recently described meta-analysis, which identified the existence of at least 4 distinct gene expression–based Consensus Molecular Subtypes (CMS), i.e. immune (CMS1), canonical (CMS2), metabolic (CMS3) and mesenchymal (CMS4) subtype. Importantly, the CMS reflect responses to presently available treatments, illustrated by the fact that only CMS2 patients respond well to standard-of-care therapy.
Cyclin A2 is an established regulator of cell proliferation and has been used for molecular diagnostics as a proliferation marker. Few studies investigated, so far, the role of cyclin A2 in tumor development in vivo using genetically altered mice.
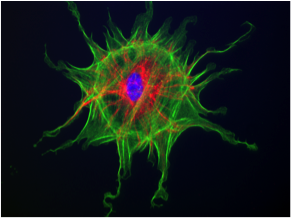
To clarify the function of cyclin A2 in colon homeostasis and colorectal cancer (CRC), we generated mice deficient for cyclin A2 in colonic epithelial cells (CEC). Colon of those mice displayed architectural changes in the mucosa, and signs of inflammation as well as an increased proliferation of CEC associated with the appearance of low-and high-grade dysplasia. The main initial events triggering those alterations in cyclin A2 deficient CEC appear to be abnormal mitoses and DNA damage. In addition, we detected an increased activity of the canonical WNT pathway, known to be deregulated in various cancers, notably CRC (Figure 1). Cyclin A2 deletion in CEC promoted the development of dysplasia and adenocarcinomas in the murine colitis-associated cancer model. We next explored the status of cyclinA2 expression in clinical CRC samples at the mRNA and protein level and found higher expression in tumors of stage I and II patients compared to those of stage III and IV. A meta-analysis of 11 transcriptome datasets comprising 2,239 primary CRC tumors displayed different CCNA2 (the mRNA coding for cyclinA2) expression levels among the CRC tumor subtypes with highest in CMS1 and lowest in CMS4, which is the most deleterious one. Moreover, high expression of CCNA2 was found to be a new independent prognosis factor for CRC tumors.
We conclude that the expression pattern of cyclin A2 in CRC tissues mirrors distinct roles during colon carcinogenesis, such as driving cell proliferation in early stages, when highly expressed, but promoting aggressiveness in later stages, when expression levels are lower. Therefore, our analysis on cyclin A2 strikingly illustrates the complexity of CRC, i.e. that a protein can execute distinct functions during tumor development complicating the development of therapeutic strategies.
The main research focus of the Hahne team is now on the role of primary cilia and related posttranslational modifications of tubulin in colonic homeostasis and pathology. For this, we employ a similar experimental approach as described for the cyclin A2 project.
Hahne team: Michael Hahne (DR2), Benedicte Lemmers (CRCN), Valerie Pinet (CRHC), Conception Paul (IEHC), Monica Gabola (PhD student), Maya Sarieddine (PhD student), Cyrine Chanchabi (AI), Vanessa Pires (AI), Asma Majoul (Master student).
TEAM GREGOIRE : Tumoral heterogeneity in hepatocellular carcinoma
Hepatocellular carcinoma (HCC) is the most common primary liver cancer and the third cause of cancer mortality worldwide. Because therapeutic options for its treatment remain very limited, there is an urgent need to better understand the disease : the implementation of new targeted therapies for hepatocellular carcinoma (HCC) requires a better characterization of the tumorigenic mechanisms resulting from the distinct driver mutations.
HCC are highly heterogenous tumours that have been classified into groups characterized by defined genomic signatures and clinico-pathological characteristics. We are interested in functional aspects of tumour heterogeneity, notably with regard to the way a genetic composition of tumoral subclones shapes the tumour microenvironment. In this context we are studying different combinations of oncogenic events that are characteristic of several classes of human HCC. Moreover, in collaboration with Daniel Fisher's team at the IGMM, we are investigating novel oncogenic functions for CDK8 and CDK19 orthologous kinases in hepatic carcinogenesis.
Our work is based on state-of-the-art mouse models of hepatic carcinogenesis. We notably use a murine model of carcinogenesis based on in vivo transfection of hepatocytes by hydrodynamic injection to generate tumors of more than 15 oncogenic combinations, on wild-type and transgenic mice. We have also developed a model based on orthotopic xenografts of cells transformed by combinations of oncogenes and labelled with fluorescent protein markers, allowing us to follow them in situ, as well as isolate and further analyse them ex vivo. These studies provide information on interactions between tumour sub-clone cell populations as well as those between the tumour and its microenvironment.
Bacevic K., Prieto S., Caruso S., Camasses A., Dubra G., Ursic-Bedoya J., Lozano A., Butterworth J., Zucman-Rossi J., Hibner U., Fisher D. *,1 and Grégoire D. *,1 (2019) The paralogue kinases CDK8 and CDK19 are essential and non-redundant in hepatic carcinogenesis. BioRxiv, 2019 789586; doi: https://doi.org/10.1101/789586
Arechederra M., Bazai S., Abdouni A., Sequera C., Mead T., Richelme S., Daian F., Audebert S., Dono R., Lozano A., Grégoire D., Hibner U., Allende D., Apte S., Maina F (2020) ADAMTSL5 is an epigenetically activated gene underlying tumorigenesis and drug resistance in hepatocellular carcinoma Journal of Hepatology. 90,6022-35. doi: https://doi.org/10.1016/j.jhep.2020.11.008
Saksouk N., Hajdari S., Perez Y., Pratlong M., Barrachina C., Graber C., Zavoriti A., Grégoire D., Sarrazin A., Pirot N., Noël JY, Khellaf L., Fabbrizio E., Julien E., Cammas F. (2020) The mouse HP1 proteins are essential for preventing liver tumorigenesis Oncogene. https://doi.org/10.1038/s41388-020-1177-8
Vegna S., Grégoire D., Moreau M., Lassus P., Durantel D., Assenat E., Hibner U., Simonin Y. (2016) NOD1 participates in the Innate Immune response triggered by Hepatitis C Virus polymerase. Journal of Virology. 90,6022-35.